
Welcome to the homepage of our bioinformatics research activities achieved in collaboration with GIGA Research teams and facilities (Interdisciplinary Cluster for Applied Genoproteomics) hosted by the University of Liège.

Here is the content of this page:
Application, analysis, and development of data mining and modeling techniques for the understanding of processes and the construction of predictive models related to complex biological processes of relevance in medical, agronomical and environmental sciences. In particular, identification of biomarkers and diagnostic/prognostic/treatment response criteria from mixed data sources (mass-spectrometry based proteomics, gene expressions, SNP based genotyping, genetic and proteomic sequences, ontologies, images).
keywords: data mining, machine learning, mass spectra, seldi, crohn, CPOD, Rheumatoide Arthritis, classification, prediction, biomarkers
In collaboration with the laboratory of clinical chemistry and rheumatology (see also project http://www.giga.ulg.ac.be/extranet/txt_databases/PROJETS/MP-I-50_Mal_Mer_Fillet.pdf),
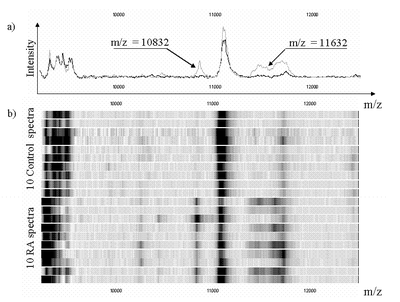
we apply machine learning techniques for the diagnosis of inflammatory diseases (Crohn, Rheumathoid Arthritis, CPOD, …) from proteomic mass spectra.
Databases containing data from healthy and disease patients have been gathered using Surface Enhanced Laser Desorption/Ionisation-Time of Flight-Mass Spectrometry (SELDI-TOF-MS). Several machine learning tools were applied to this
data. The results in terms of accuracy of the diagnostic rule and identified biomarkers are very promising.
Our results are generally superior to those obtained with standard pre- and post-processing techniques used in these applications (peak-detection and p-values based biomarker selection) and are also competitive with existing practice for the diagnosis of these diseases. The methodology is furthermore generic and it could be applied to data obtained from other medical instrumentation like for example microarray.
Send e-mail to Pierre Geurts, Raphaël Marée, Louis Wehenkel.
One of the biggest challenge of post-genomic biology is to understand and determine the complex mechanisms underlying interactions between genes and gene products. In the context of a collaboration with the AMIS-BIO team (Prof. Florence d'Alché-Buc) from the IBISC laboratory at the University of Evry, one of our research interest is to use machine learning techniques for the inference of gene interaction networks from different sources of data, experimental or not. Two kind of approaches are currently investigated:
Send e-mail to Pierre Geurts, Louis Wehenkel.
keywords: data mining, machine learning, clinico-genomic, classification, regression, cd4, prediction, biomarkers
Since October 2005, in collaboration with Prof. Michel Moutschen, this project is about the modeling of treatment response of HIV-infected patients. In such a study, there are a lot of historical patient data and the understanding of the influence of various parameters (such as age, sex, other diseases, initial CD4 level and viral load, type of tritherapy, virus mutations, ...) is not straightforward. Our approach try to combine data mining of clinical and genomic data.
This project is partially funded by a "Bourse du Standard de Liège" grant from "Léon Fredericq" foundation (2005-2007).
Send e-mail to Raphaël Marée, Louis Wehenkel.
keywords: data mining, machine learning, computer vision, image classification
With the improvements in biosensors and high-throughput image
acquisition technologies, life science laboratories are able to
perform an increasing number of experiments that involve the
generation of a large amount of images at different imaging
modalities/scales. However, manual classification
of such an amount of images is time-consuming, repetitive, and could
not always be considered reliable due to experimental conditions,
variable image quality, and human subjectivity or tiredness that lead
to considerable interobserver variations and misclassifications.
It stress the need for computer vision methods
that automate image classification tasks.
Given a set of training images labelled
into a finite number of classes, the goal of an automatic image classification
method is to build a model that will be able to predict accurately the class of
new, unseen images.
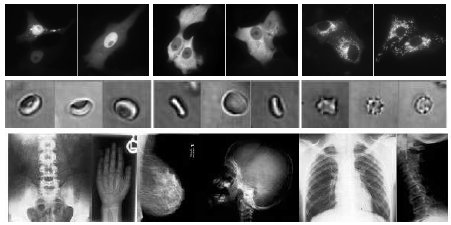
Here at the Systems and Modeling research unit, we developped and adapted a new generic Data Mining approach for image classification. It was successfully applied to several types of image classification problems. In a recent work, our method was successfully evaluated on three datasets of biological images at different imaging modalities/scales: subcellular locations, red-blood cell shapes, human body regions (X-Ray) (see publication MGW06, MGPW05b). Many biomedical applications (e.g. clinical diagnosis, subcellular location proteomics, drug discovery, ...) could benefit from this approach especially since it is directly applicable without tedious adaptation.
See related softwares: PiXiT, and Annotor.
Send e-mail to Raphaël Marée, Pierre Geurts, Louis Wehenkel.
keywords: databases, clusters, GenBank, EMBL, NCBI
We develop ad hoc search engine tools to efficiently access (collect, filter, sort, store locally) DNA sequence databases and sequence annotation databases.
These tools are developped in collaboration with Bernard Peers from Genetic and Molecular Biology Unit (CBIG), and with Sébastien Rigali from Center for Protein Engineering, University of Liège. They have motivated the development of our tools for studying genes related to the development of pancreas tissues in Zebra Fish (Danio Rerio) or to search for nucleotides that regulate the genes for a bacteria specie, but the resulting software are general and can be used in the context of similar studies.
Send e-mail to Samuel Hiard, Raphaël Marée, Louis Wehenkel.